An Integral View of the Relationship Between Genotypes and Phenotypes
Are species encoded in their genomes as molecular descriptions? Can we solve the forward problem of figuring the actions of the system from this knowledge by studying the genome, as suggested by the original concept of the ‘genetic program’?
This is both impractical and wrong, as I would contend in this article. As a result, we must abandon the gene-centric ‘differential’ interpretation of the relationship between genotype and phenotype in favor of an integrative one.

Likelihood of a successful integral view
The number of genes in the human genome is currently estimated to be about 25000 to 30000. Though the number would be much higher if we considered regions of the genome that serve as models for non-protein-coding RNAs and uncounted microRNAs. Without any additional detail, the number of possible interactions between 25000 components is approximately 1×1070 000.
Based on the number of variants and post-transcriptional changes, several more proteins are produced than genes. Proteins are the true workhorses of the body, so the estimation should be based on this figure, which could be in excess of 100000. Which are then augmented by a wide range of post-translational modifications that affect their activity.
Certainly, these calculations are unrealistic. In reality, the vast majority of conceivable interactions are impossible to realize. Because of compartmentalization, certain components never interact directly with each other, and proteins almost never interact with anything they come into contact with.
Nonetheless, we cannot rely on interaction precision to minimize the number as much as previously assumed. The majority of proteins are not very unique. Each has many interactions with other elements in the body, and many are unstructured in the sense that they lack a distinct three-dimensional structure. As a result, will shift to respond in unpredictable ways in protein and metabolic networks.
In fact, even a committed bottom-up computational biologist will search the aspects in which nature has limited the potentially feasible interactions. Organisms change in stages, with each level altering the choices available to them.
The change could be simply observed by using an antiderivative or integral calculator. Put an equation there and see how derivation reverses back with integration.
Relationships between various forms of inheritance
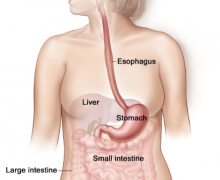
The eukaryotic cell has an astonishingly intricate structure. It is not actually a cell membrane-enclosed bag containing a protein broth. Also, prokaryotes, which were formerly considered to match the classification, are arranged and, in some cases, compartmentalized. However, the membranous organelles and other structures separate the eukaryotic cell into several more compartments.
The nucleus is strongly organized as well. Since it is more than just a container for naked DNA, nuclear fusion experiments are not stringent criteria for eliminating non-DNA inheritance.
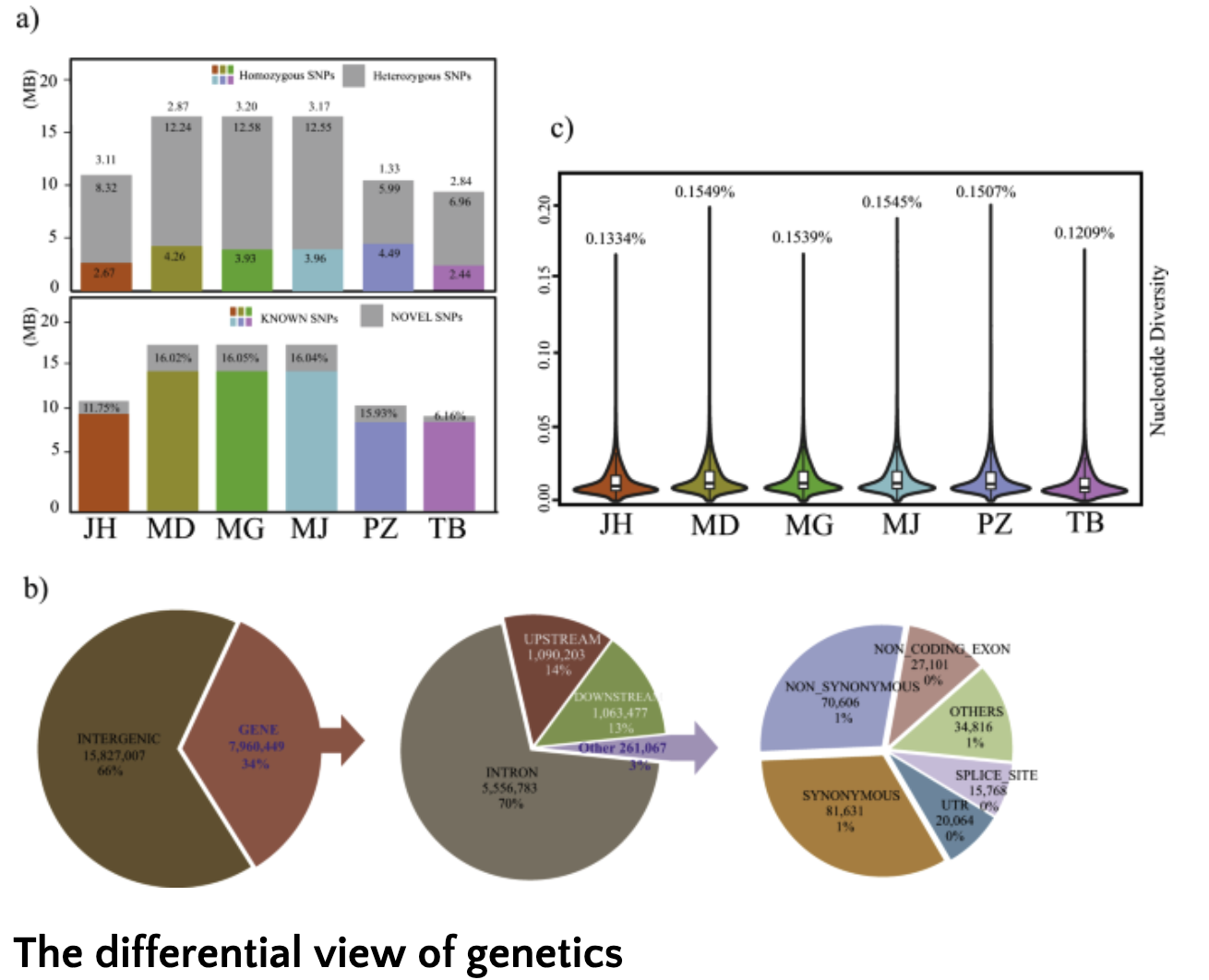
These points are self-evident and have been since electron microscopes first showed the fine details of those complex sub-cellular structures around 50 years ago. One must wonder how mainstream genetics came to neglect the problem. The solution is found in what I’ll refer to as the differential interpretation of genetics.
A brief history of genetics will be useful at this stage.
What exactly is a gene?
A gene may be characterized as something whose presence or absence affects the phenotype. When genetics and natural selection were merged to create the new synthesis known as Neo-Darwinism, the notion took root that only certain differences were important to the evolutionary success and that the only thing that mattered in applying genetics to phenotypes was to determine the genetic roots of those differences.
Since phenotype must have such an origin, selecting phenotypes is almost the same as selecting individual genes. It makes no difference in which way you look at it. They are functionally equivalent. The organism is then reduced to the position of disposable carrier of its genes, according to the gene’s-eye view.
To this point of view, we should add that in any situation, only variations in genetic make-up can be found. The technique is as simple as changing the genes by mutation, deletion, or insertion and observing the effect on the phenotype.
We must apply an implied inference to the differential viewpoint. Since, according to this viewpoint, no phenotypic variations that are not caused by a genetic variation can be inherited, the fertilized egg cell does not change other than by mutations and other means of gene evolution. Since the inherited detail in the rest of the egg cell is overlooked,
- It is believed to be identical across species. The prediction being that a cross-species clone would still exhibit the phenotype of the species that supply the genes, and
- It does not change, even if it does, it does not pass on these distinctions to future generations, which amounts to the same thing. Inheritance is required for evolution. It makes no difference if the move is just temporary.
The integral view of genetics
The integral perspective is an alternative to the differential approach. It is better described as a supplement to the differential method. We investigate a gene’s contributions to all of the processes by which its products participate.
This is the integrative biology approach, and I’m using the terms “integral” and “integrative” interchangeably. Of necessity, integrative biology does not always or inherently use mathematics, but even though it does, the comparison with mathematical integration is also valid. Because, it is not limited to examining discrepancies, and the additional knowledge taken into consideration is similar to the original and boundary conditions of mathematics.
When differential equations are used in mathematical modeling, they are precisely equivalent. The middle-out solution must be integrative. It must deal with the complications that arise as a result of taking these circumstances into account. The reason for the integrative solution is not that it makes life simpler or that it reduces uncertainty.
The other way around, uncertainty poses a significant obstacle. As a result, convincing reasons for taking this method are needed. One such point is that, in most cases, the differential method fails to expose gene functions. Many genome-level therapies, such as knockouts, are successfully buffered by the host.
Approximately 80% of knockouts are usually ‘silent’ in yeast. Although there may be underlying effects in the protein networks, these are obviously concealed by the higher levels of buffering. Indeed, one of the major and costly disappointments of recent biology has been the inability of knockouts to expose gene functions systematically and accurately.
Working only at the level of differential equations in genetics is akin to working only at the level of differential equations in algebra, as though the integral sign had never been invented. This is a strong example since the integration constants, the original and boundary conditions, restrict the potential solutions like how cell and tissue configurations restrain whatever molecular interactions are possible.
Summing Up
Finally, the integral perspective contributes to the resolution of two relevant questions in heredity and evolutionary theory. The first issue is the definition of a gene. The presence of many splice variants of several genes, as well as the probability of splicing exons from various gene sequences, has prompted some biologists to suggest that the term “gene” be redefined as the completed mRNA.
The obvious problem with this strategy is that why do we stop at the mRNA phase? Why not take it a step further and redefine the gene in terms of proteins for which DNA sequences serve as models? The difference between genotype and phenotype will become less apparent and could even vanish.